
RNSA 2023 Abdominal Trauma Detection
Install Libraries¶
#Install Libraries
!python -m pip install --no-index --find-links=/kaggle/input/libraries seaborn
!python -m pip install --no-index --find-links=/kaggle/input/libraries keras-cv-attention-models
!python -m pip install --no-index --find-links=/kaggle/input/libraries scikit-learn
#!python -m pip install --no-index --find-links=/kaggle/input/libraries-add02 python-gdcm
!python -m pip install --no-index --find-links=/kaggle/input/libraries-add02 pylibjpeg
!python -m pip install --no-index --find-links=/kaggle/input/libraries-add02 pydicom
Looking in links: /kaggle/input/libraries Requirement already satisfied: seaborn in /opt/conda/lib/python3.10/site-packages (0.12.2) Requirement already satisfied: numpy!=1.24.0,>=1.17 in /opt/conda/lib/python3.10/site-packages (from seaborn) (1.23.5) Requirement already satisfied: pandas>=0.25 in /opt/conda/lib/python3.10/site-packages (from seaborn) (2.0.2) Requirement already satisfied: matplotlib!=3.6.1,>=3.1 in /opt/conda/lib/python3.10/site-packages (from seaborn) (3.7.2) Requirement already satisfied: contourpy>=1.0.1 in /opt/conda/lib/python3.10/site-packages (from matplotlib!=3.6.1,>=3.1->seaborn) (1.1.0) Requirement already satisfied: cycler>=0.10 in /opt/conda/lib/python3.10/site-packages (from matplotlib!=3.6.1,>=3.1->seaborn) (0.11.0) Requirement already satisfied: fonttools>=4.22.0 in /opt/conda/lib/python3.10/site-packages (from matplotlib!=3.6.1,>=3.1->seaborn) (4.40.0) Requirement already satisfied: kiwisolver>=1.0.1 in /opt/conda/lib/python3.10/site-packages (from matplotlib!=3.6.1,>=3.1->seaborn) (1.4.4) Requirement already satisfied: packaging>=20.0 in /opt/conda/lib/python3.10/site-packages (from matplotlib!=3.6.1,>=3.1->seaborn) (21.3) Requirement already satisfied: pillow>=6.2.0 in /opt/conda/lib/python3.10/site-packages (from matplotlib!=3.6.1,>=3.1->seaborn) (9.5.0) Requirement already satisfied: pyparsing<3.1,>=2.3.1 in /opt/conda/lib/python3.10/site-packages (from matplotlib!=3.6.1,>=3.1->seaborn) (3.0.9) Requirement already satisfied: python-dateutil>=2.7 in /opt/conda/lib/python3.10/site-packages (from matplotlib!=3.6.1,>=3.1->seaborn) (2.8.2) Requirement already satisfied: pytz>=2020.1 in /opt/conda/lib/python3.10/site-packages (from pandas>=0.25->seaborn) (2023.3) Requirement already satisfied: tzdata>=2022.1 in /opt/conda/lib/python3.10/site-packages (from pandas>=0.25->seaborn) (2023.3) Requirement already satisfied: six>=1.5 in /opt/conda/lib/python3.10/site-packages (from python-dateutil>=2.7->matplotlib!=3.6.1,>=3.1->seaborn) (1.16.0) Looking in links: /kaggle/input/libraries Processing /kaggle/input/libraries/keras_cv_attention_models-1.3.20-py3-none-any.whl Requirement already satisfied: pillow in /opt/conda/lib/python3.10/site-packages (from keras-cv-attention-models) (9.5.0) Processing /kaggle/input/libraries/ftfy-6.1.1-py3-none-any.whl (from keras-cv-attention-models) Requirement already satisfied: regex in /opt/conda/lib/python3.10/site-packages (from keras-cv-attention-models) (2023.6.3) Requirement already satisfied: tensorflow-datasets in /opt/conda/lib/python3.10/site-packages (from keras-cv-attention-models) (4.9.2) Requirement already satisfied: tensorflow in /opt/conda/lib/python3.10/site-packages (from keras-cv-attention-models) (2.12.0) Requirement already satisfied: wcwidth>=0.2.5 in /opt/conda/lib/python3.10/site-packages (from ftfy->keras-cv-attention-models) (0.2.6) Requirement already satisfied: absl-py>=1.0.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (1.4.0) Requirement already satisfied: astunparse>=1.6.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (1.6.3) Requirement already satisfied: flatbuffers>=2.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (23.5.26) Requirement already satisfied: gast<=0.4.0,>=0.2.1 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (0.4.0) Requirement already satisfied: google-pasta>=0.1.1 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (0.2.0) Requirement already satisfied: grpcio<2.0,>=1.24.3 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (1.51.1) Requirement already satisfied: h5py>=2.9.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (3.9.0) Requirement already satisfied: jax>=0.3.15 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (0.4.13) Requirement already satisfied: keras<2.13,>=2.12.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (2.12.0) Requirement already satisfied: libclang>=13.0.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (16.0.0) Requirement already satisfied: numpy<1.24,>=1.22 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (1.23.5) Requirement already satisfied: opt-einsum>=2.3.2 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (3.3.0) Requirement already satisfied: packaging in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (21.3) Requirement already satisfied: protobuf!=4.21.0,!=4.21.1,!=4.21.2,!=4.21.3,!=4.21.4,!=4.21.5,<5.0.0dev,>=3.20.3 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (3.20.3) Requirement already satisfied: setuptools in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (68.0.0) Requirement already satisfied: six>=1.12.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (1.16.0) Requirement already satisfied: tensorboard<2.13,>=2.12 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (2.12.3) Requirement already satisfied: tensorflow-estimator<2.13,>=2.12.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (2.12.0) Requirement already satisfied: termcolor>=1.1.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (2.3.0) Requirement already satisfied: typing-extensions>=3.6.6 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (4.6.3) Requirement already satisfied: wrapt<1.15,>=1.11.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (1.14.1) Requirement already satisfied: tensorflow-io-gcs-filesystem>=0.23.1 in /opt/conda/lib/python3.10/site-packages (from tensorflow->keras-cv-attention-models) (0.32.0) Requirement already satisfied: array-record in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (0.4.0) Requirement already satisfied: click in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (8.1.7) Requirement already satisfied: dm-tree in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (0.1.8) Requirement already satisfied: etils[enp,epath]>=0.9.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (1.3.0) Requirement already satisfied: promise in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (2.3) Requirement already satisfied: psutil in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (5.9.3) Requirement already satisfied: requests>=2.19.0 in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (2.31.0) Requirement already satisfied: tensorflow-metadata in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (0.14.0) Requirement already satisfied: toml in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (0.10.2) Requirement already satisfied: tqdm in /opt/conda/lib/python3.10/site-packages (from tensorflow-datasets->keras-cv-attention-models) (4.66.1) Requirement already satisfied: wheel<1.0,>=0.23.0 in /opt/conda/lib/python3.10/site-packages (from astunparse>=1.6.0->tensorflow->keras-cv-attention-models) (0.40.0) Requirement already satisfied: importlib_resources in /opt/conda/lib/python3.10/site-packages (from etils[enp,epath]>=0.9.0->tensorflow-datasets->keras-cv-attention-models) (5.12.0) Requirement already satisfied: zipp in /opt/conda/lib/python3.10/site-packages (from etils[enp,epath]>=0.9.0->tensorflow-datasets->keras-cv-attention-models) (3.15.0) Requirement already satisfied: ml-dtypes>=0.1.0 in /opt/conda/lib/python3.10/site-packages (from jax>=0.3.15->tensorflow->keras-cv-attention-models) (0.2.0) Requirement already satisfied: scipy>=1.7 in /opt/conda/lib/python3.10/site-packages (from jax>=0.3.15->tensorflow->keras-cv-attention-models) (1.11.2) Requirement already satisfied: charset-normalizer<4,>=2 in /opt/conda/lib/python3.10/site-packages (from requests>=2.19.0->tensorflow-datasets->keras-cv-attention-models) (3.1.0) Requirement already satisfied: idna<4,>=2.5 in /opt/conda/lib/python3.10/site-packages (from requests>=2.19.0->tensorflow-datasets->keras-cv-attention-models) (3.4) Requirement already satisfied: urllib3<3,>=1.21.1 in /opt/conda/lib/python3.10/site-packages (from requests>=2.19.0->tensorflow-datasets->keras-cv-attention-models) (1.26.15) Requirement already satisfied: certifi>=2017.4.17 in /opt/conda/lib/python3.10/site-packages (from requests>=2.19.0->tensorflow-datasets->keras-cv-attention-models) (2023.7.22) Requirement already satisfied: google-auth<3,>=1.6.3 in /opt/conda/lib/python3.10/site-packages (from tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (2.20.0) Requirement already satisfied: google-auth-oauthlib<1.1,>=0.5 in /opt/conda/lib/python3.10/site-packages (from tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (1.0.0) Requirement already satisfied: markdown>=2.6.8 in /opt/conda/lib/python3.10/site-packages (from tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (3.4.3) Requirement already satisfied: tensorboard-data-server<0.8.0,>=0.7.0 in /opt/conda/lib/python3.10/site-packages (from tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (0.7.1) Requirement already satisfied: werkzeug>=1.0.1 in /opt/conda/lib/python3.10/site-packages (from tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (2.3.7) Requirement already satisfied: pyparsing!=3.0.5,>=2.0.2 in /opt/conda/lib/python3.10/site-packages (from packaging->tensorflow->keras-cv-attention-models) (3.0.9) Requirement already satisfied: googleapis-common-protos in /opt/conda/lib/python3.10/site-packages (from tensorflow-metadata->tensorflow-datasets->keras-cv-attention-models) (1.59.1) Requirement already satisfied: cachetools<6.0,>=2.0.0 in /opt/conda/lib/python3.10/site-packages (from google-auth<3,>=1.6.3->tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (4.2.4) Requirement already satisfied: pyasn1-modules>=0.2.1 in /opt/conda/lib/python3.10/site-packages (from google-auth<3,>=1.6.3->tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (0.2.7) Requirement already satisfied: rsa<5,>=3.1.4 in /opt/conda/lib/python3.10/site-packages (from google-auth<3,>=1.6.3->tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (4.9) Requirement already satisfied: requests-oauthlib>=0.7.0 in /opt/conda/lib/python3.10/site-packages (from google-auth-oauthlib<1.1,>=0.5->tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (1.3.1) Requirement already satisfied: MarkupSafe>=2.1.1 in /opt/conda/lib/python3.10/site-packages (from werkzeug>=1.0.1->tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (2.1.3) Requirement already satisfied: pyasn1<0.5.0,>=0.4.6 in /opt/conda/lib/python3.10/site-packages (from pyasn1-modules>=0.2.1->google-auth<3,>=1.6.3->tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (0.4.8) Requirement already satisfied: oauthlib>=3.0.0 in /opt/conda/lib/python3.10/site-packages (from requests-oauthlib>=0.7.0->google-auth-oauthlib<1.1,>=0.5->tensorboard<2.13,>=2.12->tensorflow->keras-cv-attention-models) (3.2.2) Installing collected packages: ftfy, keras-cv-attention-models Successfully installed ftfy-6.1.1 keras-cv-attention-models-1.3.20 Looking in links: /kaggle/input/libraries Requirement already satisfied: scikit-learn in /opt/conda/lib/python3.10/site-packages (1.2.2) Requirement already satisfied: numpy>=1.17.3 in /opt/conda/lib/python3.10/site-packages (from scikit-learn) (1.23.5) Requirement already satisfied: scipy>=1.3.2 in /opt/conda/lib/python3.10/site-packages (from scikit-learn) (1.11.2) Requirement already satisfied: joblib>=1.1.1 in /opt/conda/lib/python3.10/site-packages (from scikit-learn) (1.3.2) Requirement already satisfied: threadpoolctl>=2.0.0 in /opt/conda/lib/python3.10/site-packages (from scikit-learn) (3.1.0) Looking in links: /kaggle/input/libraries-add02 Processing /kaggle/input/libraries-add02/pylibjpeg-1.4.0-py3-none-any.whl Requirement already satisfied: numpy in /opt/conda/lib/python3.10/site-packages (from pylibjpeg) (1.23.5) Installing collected packages: pylibjpeg Successfully installed pylibjpeg-1.4.0 Looking in links: /kaggle/input/libraries-add02 Requirement already satisfied: pydicom in /opt/conda/lib/python3.10/site-packages (2.4.3)
Import Libraries¶
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3' # to avoid too many logging messages
import pandas as pd, numpy as np, random, shutil
import tensorflow as tf, re, math
import tensorflow.keras.backend as K
import sklearn
import matplotlib.pyplot as plt
import tensorflow_addons as tfa
import tensorflow_probability as tfp
import yaml
from IPython import display as ipd
from glob import glob
from tqdm import tqdm
from sklearn.model_selection import KFold, StratifiedKFold, GroupKFold, StratifiedGroupKFold
from sklearn.metrics import roc_auc_score
from sklearn.utils.class_weight import compute_class_weight
import time
import pydicom
/opt/conda/lib/python3.10/site-packages/scipy/__init__.py:146: UserWarning: A NumPy version >=1.16.5 and <1.23.0 is required for this version of SciPy (detected version 1.23.5
warnings.warn(f"A NumPy version >={np_minversion} and <{np_maxversion}"
/opt/conda/lib/python3.10/site-packages/tensorflow_addons/utils/tfa_eol_msg.py:23: UserWarning:
TensorFlow Addons (TFA) has ended development and introduction of new features.
TFA has entered a minimal maintenance and release mode until a planned end of life in May 2024.
Please modify downstream libraries to take dependencies from other repositories in our TensorFlow community (e.g. Keras, Keras-CV, and Keras-NLP).
For more information see: https://github.com/tensorflow/addons/issues/2807
warnings.warn()
class CFG:
competition = 'rsna-atd'
debug = False
comment = 'EfficientNetV1B0-256x256-low_lr-vflip'
exp_name = 'baseline-v4: new_ds + multi_head' # name of the experiment, folds will be grouped using 'exp_name'
verbose = 0
display_plot = True
# device
device = "GPU" #or "TPU-VM"
model_name = 'EfficientNetV1B0'
# seed for data-split, layer init, augs
seed = 42
# number of folds for data-split
folds = 4
# which folds to train
selected_folds = [0, 1, 2]
# size of the image
img_size = [256, 256]
# dicom to png size
resize_dim = 512
# batch_size and epochs
batch_size = 16
epochs = 10
# loss
loss = 'BCE & CCE' # BCE, Focal
# optimizer
optimizer = 'Adam'
# augmentation
augment = True
# scale-shift-rotate-shear
transform = 0.90 # transform prob
fill_mode = 'constant'
rot = 2.0
shr = 2.0
hzoom = 50.0
wzoom = 50.0
hshift = 10.0
wshift = 10.0
# flip
hflip = True
vflip = True
# clip
clip = False
# lr-scheduler
scheduler = 'cosine' # cosine
# dropout
drop_prob = 0.6
drop_cnt = 5
drop_size = 0.05
# cut-mix-up
mixup_prob = 0.0
mixup_alpha = 0.5
cutmix_prob = 0.0
cutmix_alpha = 2.5
# pixel-augment
pixel_aug = 0.90 # prob of pixel_aug
sat = [0.7, 1.3]
cont = [0.8, 1.2]
bri = 0.15
hue = 0.05
# test-time augs
tta = 1
# target column
target_col = [ "bowel_injury", "extravasation_injury", "kidney_healthy", "kidney_low",
"kidney_high", "liver_healthy", "liver_low", "liver_high",
"spleen_healthy", "spleen_low", "spleen_high"] # not using "bowel_healthy" & "extravasation_healthy"
Reproducibility¶
def seeding(SEED):
np.random.seed(SEED)
random.seed(SEED)
os.environ['PYTHONHASHSEED'] = str(SEED)
tf.random.set_seed(SEED)
print('seeding done!!!')
seeding(CFG.seed)
seeding done!!!
Device Configs¶
This notebook is compatible for remote-tpu, local-tpu, multi-gpu and single-gpu. Simple change to device="TPU" for remote-tpu and device="TPU-VM" for local-tpu and finally, device="GPU" for single or multi-gpu.
if "TPU" in CFG.device:
tpu = 'local' if CFG.device=='TPU-VM' else None
print("connecting to TPU...")
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver.connect(tpu=tpu)
strategy = tf.distribute.TPUStrategy(tpu)
except:
CFG.device = "GPU"
if CFG.device == "GPU" or CFG.device=="CPU":
ngpu = len(tf.config.experimental.list_physical_devices('GPU'))
if ngpu>1:
print("Using multi GPU")
strategy = tf.distribute.MirroredStrategy()
elif ngpu==1:
print("Using single GPU")
strategy = tf.distribute.get_strategy()
else:
print("Using CPU")
strategy = tf.distribute.get_strategy()
CFG.device = "CPU"
if CFG.device == "GPU":
print("Num GPUs Available: ", ngpu)
AUTO = tf.data.experimental.AUTOTUNE
REPLICAS = strategy.num_replicas_in_sync
print(f'REPLICAS: {REPLICAS}')
Using single GPU
Num GPUs Available: 1
REPLICAS: 1
ROOT PATH¶
BASE_PATH = f'/kaggle/input/rsna-atd-512x512-png-v2-dataset'
# train
df = pd.read_csv(f'{BASE_PATH}/train.csv')
df['image_path'] = f'{BASE_PATH}/train_images'\
+ '/' + df.patient_id.astype(str)\
+ '/' + df.series_id.astype(str)\
+ '/' + df.instance_number.astype(str) +'.png'
df = df.drop_duplicates()
print('Train:')
display(df.head(2))
# test
test_df = pd.read_csv(f'{BASE_PATH}/test.csv')
test_df['image_path'] = f'{BASE_PATH}/test_images'\
+ '/' + test_df.patient_id.astype(str)\
+ '/' + test_df.series_id.astype(str)\
+ '/' + test_df.instance_number.astype(str) +'.png'
test_df = test_df.drop_duplicates()
print('\nTest:')
display(test_df.head(2))
Train:
| patient_id | bowel_healthy | bowel_injury | extravasation_healthy | extravasation_injury | kidney_healthy | kidney_low | kidney_high | liver_healthy | liver_low | ... | spleen_healthy | spleen_low | spleen_high | any_injury | series_id | instance_number | injury_name | image_path | width | height | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 10004 | 1 | 0 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | ... | 0 | 0 | 1 | 1 | 21057 | 362 | Active_Extravasation | /kaggle/input/rsna-atd-512x512-png-v2-dataset/... | 512 | 512 |
| 1 | 10004 | 1 | 0 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | ... | 0 | 0 | 1 | 1 | 21057 | 363 | Active_Extravasation | /kaggle/input/rsna-atd-512x512-png-v2-dataset/... | 512 | 512 |
2 rows × 21 columns
Test:
| image_path | patient_id | series_id | instance_number | width | height | |
|---|---|---|---|---|---|---|
| 0 | /kaggle/input/rsna-atd-512x512-png-v2-dataset/... | 63706 | 39279 | 30 | 512 | 512 |
| 1 | /kaggle/input/rsna-atd-512x512-png-v2-dataset/... | 50046 | 24574 | 30 | 512 | 512 |
Check If Data Exist?¶
tf.io.gfile.exists(df.image_path.iloc[0]), tf.io.gfile.exists(test_df.image_path.iloc[0])
(True, True)
Train-Test Ditribution¶
print('train_files:',df.shape[0])
print('test_files:',test_df.shape[0])
train_files: 12029
test_files: 3
df['stratify'] = ''
for col in CFG.target_col:
df['stratify'] += df[col].astype(str)
df = df.reset_index(drop=True)
skf = StratifiedGroupKFold(n_splits=CFG.folds, shuffle=True, random_state=CFG.seed)
for fold, (train_idx, val_idx) in enumerate(skf.split(df, df['stratify'], df["patient_id"])):
df.loc[val_idx, 'fold'] = fold
display(df.groupby(['fold', 'patient_id']).size())
/opt/conda/lib/python3.10/site-packages/sklearn/model_selection/_split.py:909: UserWarning: The least populated class in y has only 1 members, which is less than n_splits=4.
warnings.warn(
fold patient_id
0.0 43 53
263 12
2602 17
4331 27
5337 2
..
3.0 60993 29
62179 21
63665 71
64091 9
65456 16
Length: 246, dtype: int64
Data Augmentation¶
Used simple augmentations, some of them may hurt the model.
- RandomFlip (Left-Right)
- No Rotation
- RandomBrightness
- RndomContrast
- Shear
- Zoom
- Coarsee Dropout/Cutout
def get_mat(shear, height_zoom, width_zoom, height_shift, width_shift):
# returns 3x3 transformmatrix which transforms indicies
# CONVERT DEGREES TO RADIANS
shear = math.pi * shear / 180.
def get_3x3_mat(lst):
return tf.reshape(tf.concat([lst],axis=0), [3,3])
# ROTATION MATRIX
one = tf.constant([1],dtype='float32')
zero = tf.constant([0],dtype='float32')
# SHEAR MATRIX
c2 = tf.math.cos(shear)
s2 = tf.math.sin(shear)
shear_matrix = get_3x3_mat([one, s2, zero,
zero, c2, zero,
zero, zero, one])
# ZOOM MATRIX
zoom_matrix = get_3x3_mat([one/height_zoom, zero, zero,
zero, one/width_zoom, zero,
zero, zero, one])
# SHIFT MATRIX
shift_matrix = get_3x3_mat([one, zero, height_shift,
zero, one, width_shift,
zero, zero, one])
return K.dot(shear_matrix,K.dot(zoom_matrix, shift_matrix)) #K.dot(K.dot(rotation_matrix, shear_matrix), K.dot(zoom_matrix, shift_matrix))
def transform(image, DIM=CFG.img_size):#[rot,shr,h_zoom,w_zoom,h_shift,w_shift]):
if DIM[0]>DIM[1]:
diff = (DIM[0]-DIM[1])
pad = [diff//2, diff//2 + diff%2]
image = tf.pad(image, [[0, 0], [pad[0], pad[1]],[0, 0]])
NEW_DIM = DIM[0]
elif DIM[0]<DIM[1]:
diff = (DIM[1]-DIM[0])
pad = [diff//2, diff//2 + diff%2]
image = tf.pad(image, [[pad[0], pad[1]], [0, 0],[0, 0]])
NEW_DIM = DIM[1]
rot = CFG.rot * tf.random.normal([1], dtype='float32')
shr = CFG.shr * tf.random.normal([1], dtype='float32')
h_zoom = 1.0 + tf.random.normal([1], dtype='float32') / CFG.hzoom
w_zoom = 1.0 + tf.random.normal([1], dtype='float32') / CFG.wzoom
h_shift = CFG.hshift * tf.random.normal([1], dtype='float32')
w_shift = CFG.wshift * tf.random.normal([1], dtype='float32')
transformation_matrix=tf.linalg.inv(get_mat(shr,h_zoom,w_zoom,h_shift,w_shift))
flat_tensor=tfa.image.transform_ops.matrices_to_flat_transforms(transformation_matrix)
image=tfa.image.transform(image,flat_tensor, fill_mode=CFG.fill_mode)
rotation = math.pi * rot / 180.
image=tfa.image.rotate(image,-rotation, fill_mode=CFG.fill_mode)
if DIM[0]>DIM[1]:
image=tf.reshape(image, [NEW_DIM, NEW_DIM,3])
image = image[:, pad[0]:-pad[1],:]
elif DIM[1]>DIM[0]:
image=tf.reshape(image, [NEW_DIM, NEW_DIM,3])
image = image[pad[0]:-pad[1],:,:]
image = tf.reshape(image, [*DIM, 3])
return image
def dropout(image,DIM=CFG.img_size, PROBABILITY = 0.6, CT = 5, SZ = 0.1):
# input image - is one image of size [dim,dim,3] not a batch of [b,dim,dim,3]
# output - image with CT squares of side size SZ*DIM removed
# DO DROPOUT WITH PROBABILITY DEFINED ABOVE
P = tf.cast( tf.random.uniform([],0,1)<PROBABILITY, tf.int32)
if (P==0)|(CT==0)|(SZ==0):
return image
for k in range(CT):
# CHOOSE RANDOM LOCATION
x = tf.cast( tf.random.uniform([],0,DIM[1]),tf.int32)
y = tf.cast( tf.random.uniform([],0,DIM[0]),tf.int32)
# COMPUTE SQUARE
WIDTH = tf.cast( SZ*min(DIM),tf.int32) * P
ya = tf.math.maximum(0,y-WIDTH//2)
yb = tf.math.minimum(DIM[0],y+WIDTH//2)
xa = tf.math.maximum(0,x-WIDTH//2)
xb = tf.math.minimum(DIM[1],x+WIDTH//2)
# DROPOUT IMAGE
one = image[ya:yb,0:xa,:]
two = tf.zeros([yb-ya,xb-xa,3], dtype = image.dtype)
three = image[ya:yb,xb:DIM[1],:]
middle = tf.concat([one,two,three],axis=1)
image = tf.concat([image[0:ya,:,:],middle,image[yb:DIM[0],:,:]],axis=0)
image = tf.reshape(image,[*DIM,3])
return image
CutMixUp¶
def random_int(shape=[], minval=0, maxval=1):
return tf.random.uniform(
shape=shape, minval=minval, maxval=maxval, dtype=tf.int32)
def random_float(shape=[], minval=0.0, maxval=1.0):
rnd = tf.random.uniform(
shape=shape, minval=minval, maxval=maxval, dtype=tf.float32)
return rnd
# mixup
def get_mixup(alpha=0.2, prob=0.5):
@tf.function
def mixup(images, labels, alpha=alpha, prob=prob):
if random_float() > prob:
return images, labels
image_shape = tf.shape(images)
label_shape = tf.shape(labels)
beta = tfp.distributions.Beta(alpha, alpha)
lam = beta.sample(1)[0]
images = lam * images + (1.0 - lam) * tf.roll(images, shift=1, axis=0)
labels = lam * labels + (1.0 - lam) * tf.roll(labels, shift=1, axis=0)
images = tf.reshape(images, image_shape)
labels = tf.reshape(labels, label_shape)
return images, labels
return mixup
# cutmix
def get_cutmix(alpha, prob=0.5):
@tf.function
def cutmix(images, labels, alpha=alpha, prob=prob):
if random_float() > prob:
return images, labels
image_shape = tf.shape(images)
label_shape = tf.shape(labels)
W = tf.cast(image_shape[2], tf.int32)
H = tf.cast(image_shape[1], tf.int32)
beta = tfp.distributions.Beta(alpha, alpha)
lam = beta.sample(1)[0]
images_rolled = tf.roll(images, shift=1, axis=0)
labels_rolled = tf.roll(labels, shift=1, axis=0)
r_x = random_int([], minval=0, maxval=W)
r_y = random_int([], minval=0, maxval=H)
r = 0.5 * tf.math.sqrt(1.0 - lam)
r_w_half = tf.cast(r * tf.cast(W, tf.float32), tf.int32)
r_h_half = tf.cast(r * tf.cast(H, tf.float32), tf.int32)
x1 = tf.cast(tf.clip_by_value(r_x - r_w_half, 0, W), tf.int32)
x2 = tf.cast(tf.clip_by_value(r_x + r_w_half, 0, W), tf.int32)
y1 = tf.cast(tf.clip_by_value(r_y - r_h_half, 0, H), tf.int32)
y2 = tf.cast(tf.clip_by_value(r_y + r_h_half, 0, H), tf.int32)
# outer-pad patch -> [0, 0, 1, 1, 0, 0]
patch1 = images[:, y1:y2, x1:x2, :] # [batch, height, width, channel]
patch1 = tf.pad(
patch1, [[0, 0], [y1, H - y2], [x1, W - x2], [0, 0]]) # outer-pad
# inner-pad patch -> [1, 1, 0, 0, 1, 1]
patch2 = images_rolled[:, y1:y2, x1:x2, :]
patch2 = tf.pad(
patch2, [[0, 0], [y1, H - y2], [x1, W - x2], [0, 0]]) # outer-pad
patch2 = images_rolled - patch2 # inner-pad = img - outer-pad
images = patch1 + patch2 # cutmix img
lam = tf.cast((1.0 - (x2 - x1) * (y2 - y1) / (W * H)), tf.float32) # no H as (y1 - y2)/H = 1
labels = lam * labels + (1.0 - lam) * labels_rolled # cutmix label
images = tf.reshape(images, image_shape)
labels = tf.reshape(labels, label_shape)
return images, labels
return cutmix
Data Pipeline¶
- Reads the raw file and then decodes it to tf.Tensor
- Resizes the image in desired size
- Chages the datatype to float32
- Caches the Data for boosting up the speed.
- Uses Augmentations to reduce overfitting and make model more robust.
- Finally, splits the data into batches.
def build_decoder(with_labels=True, target_size=CFG.img_size, ext='png'):
def decode_image(path):
file_bytes = tf.io.read_file(path)
if ext == 'png':
img = tf.image.decode_png(file_bytes, channels=3, dtype=tf.uint8)
elif ext in ['jpg', 'jpeg']:
img = tf.image.decode_jpeg(file_bytes, channels=3)
else:
raise ValueError("Image extension not supported")
img = tf.image.resize(img, target_size, method='bilinear')
img = tf.cast(img, tf.float32) / 255.0
img = tf.reshape(img, [*target_size, 3])
return img
def decode_label(label):
label = tf.cast(label, tf.float32)
return (label[0:1], label[1:2], label[2:5], label[5:8], label[8:11])
def decode_with_labels(path, label):
return decode_image(path), decode_label(label)
return decode_with_labels if with_labels else decode_image
def build_augmenter(with_labels=True, dim=CFG.img_size):
def augment(img, dim=dim):
if random_float() < CFG.transform:
img = transform(img,DIM=dim)
img = tf.image.random_flip_left_right(img) if CFG.hflip else img
img = tf.image.random_flip_up_down(img) if CFG.vflip else img
if random_float() < CFG.pixel_aug:
img = tf.image.random_hue(img, CFG.hue)
img = tf.image.random_saturation(img, CFG.sat[0], CFG.sat[1])
img = tf.image.random_contrast(img, CFG.cont[0], CFG.cont[1])
img = tf.image.random_brightness(img, CFG.bri)
img = tf.clip_by_value(img, 0, 1) if CFG.clip else img
img = tf.reshape(img, [*dim, 3])
return img
def augment_with_labels(img, label):
return augment(img), label
return augment_with_labels if with_labels else augment
def build_dataset(paths, labels=None, batch_size=32, cache=True,
decode_fn=None, augment_fn=None,
augment=True, repeat=True, shuffle=1024,
cache_dir="", drop_remainder=False):
if cache_dir != "" and cache is True:
os.makedirs(cache_dir, exist_ok=True)
if decode_fn is None:
decode_fn = build_decoder(labels is not None)
if augment_fn is None:
augment_fn = build_augmenter(labels is not None)
AUTO = tf.data.experimental.AUTOTUNE
slices = paths if labels is None else (paths, labels)
ds = tf.data.Dataset.from_tensor_slices(slices)
ds = ds.map(decode_fn, num_parallel_calls=AUTO)
ds = ds.cache(cache_dir) if cache else ds
ds = ds.repeat() if repeat else ds
if shuffle:
ds = ds.shuffle(shuffle, seed=CFG.seed)
opt = tf.data.Options()
opt.experimental_deterministic = False
ds = ds.with_options(opt)
ds = ds.map(augment_fn, num_parallel_calls=AUTO) if augment else ds
if augment and labels is not None:
ds = ds.map(lambda img, label: (dropout(img,
DIM=CFG.img_size,
PROBABILITY=CFG.drop_prob,
CT=CFG.drop_cnt,
SZ=CFG.drop_size), label),num_parallel_calls=AUTO)
ds = ds.batch(batch_size, drop_remainder=drop_remainder)
if augment and labels is not None:
if CFG.cutmix_prob:
ds = ds.map(get_cutmix(alpha=CFG.cutmix_alpha,prob=CFG.cutmix_prob),num_parallel_calls=AUTO)
if CFG.mixup_prob:
ds = ds.map(get_mixup(alpha=CFG.mixup_alpha,prob=CFG.mixup_prob),num_parallel_calls=AUTO)
ds = ds.prefetch(AUTO)
return ds
Visualization¶
- Check if augmentation is working properly or not.
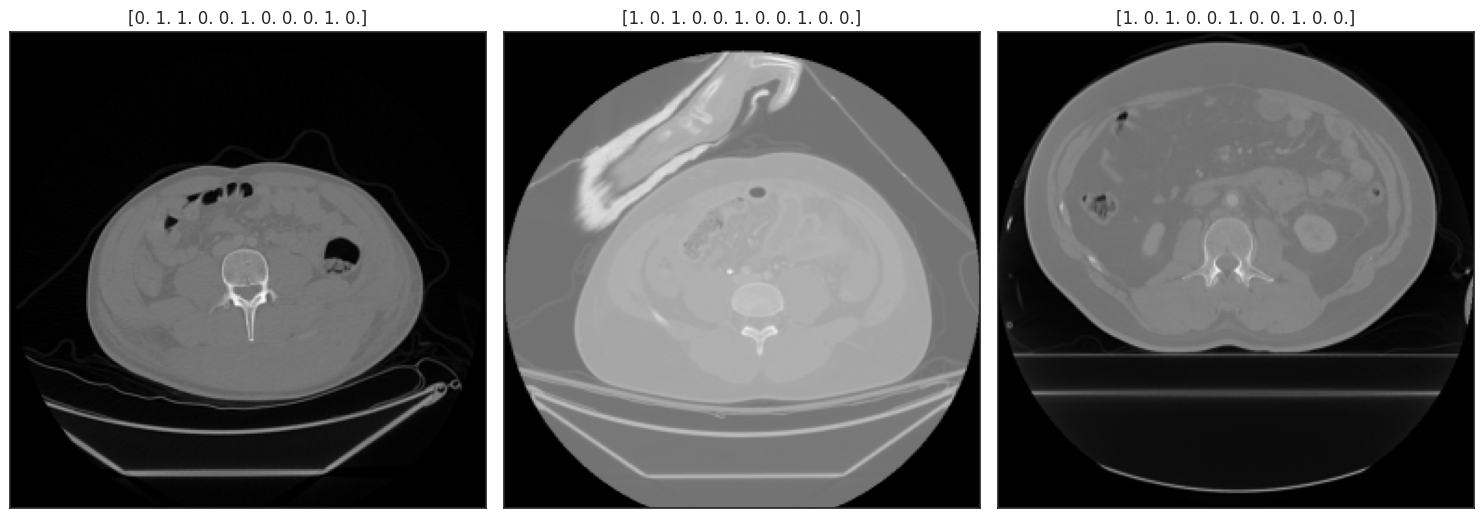
def display_batch(batch, size=2):
if isinstance(batch, tuple):
imgs, tars = batch
else:
imgs = batch
tars = None
tars = tf.concat(tars,axis=-1).numpy()
plt.figure(figsize=(size*5, 10))
for img_idx in range(size):
plt.subplot(1, size, img_idx+1)
if tars is not None:
plt.title(f'{tars[img_idx].round(2)}', fontsize=12)
img = imgs[img_idx,]
plt.imshow(img)
plt.xticks([]); plt.yticks([])
plt.tight_layout()
plt.show()
Generate Image¶
fold = 0
fold_df = df[df.fold==fold].sample(frac=1.0)
paths = fold_df.image_path.tolist()
labels = fold_df[CFG.target_col].values
ds = build_dataset(paths, labels, cache=False, batch_size=32,
repeat=True, shuffle=True, augment=False)
ds = ds.unbatch().batch(20)
batch = next(iter(ds))
No-Augmentation¶
display_batch(batch, 3);
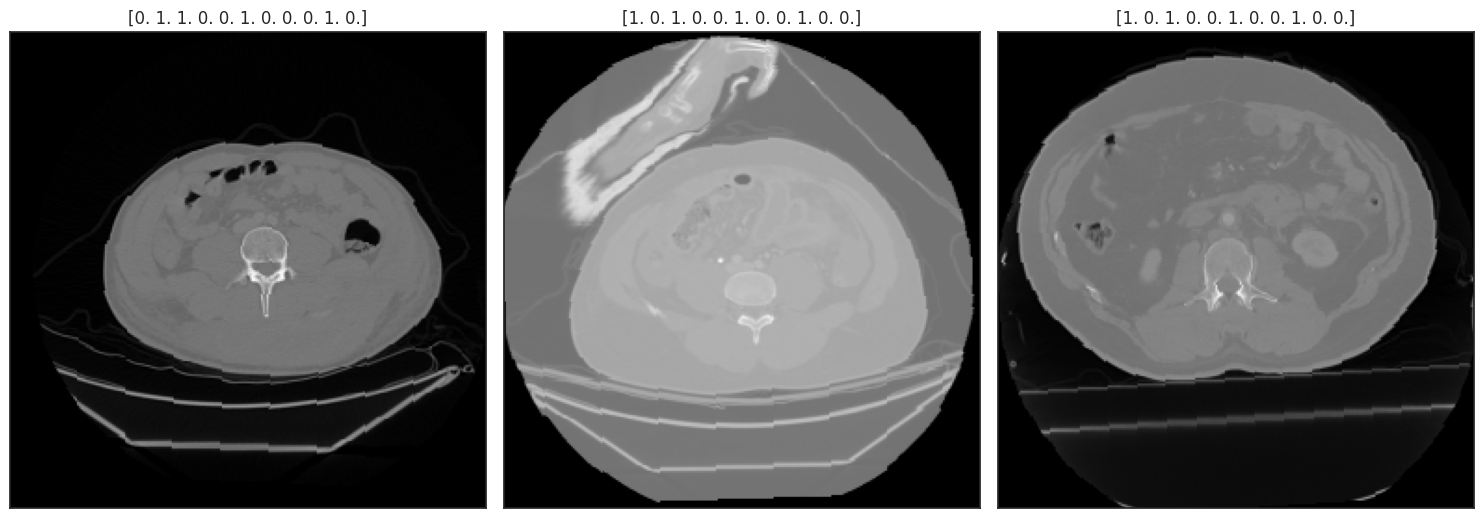
Dropout¶
dimgs = tf.map_fn(lambda img: dropout(img,
DIM=CFG.img_size,
PROBABILITY=1.0,
CT=10,
SZ=0.08), batch[0])
dtars = batch[1]
display_batch((dimgs, dtars), 3);

Affine Transform¶
timgs = tf.map_fn(lambda img: transform(img,DIM=CFG.img_size), batch[0])
ttars = batch[1]
display_batch((timgs, ttars), 3);
Loss Function¶
We'll use Binary Cross Entropy loss for bowel and extravasation at the same time we'll use Categorical Cross Entropy for kidney, liver and spleen. <!-- ## BCE Loss Function for this notebook is BCE: Binary Crossentropy or Focal as the task is binary classification
where $\hat{y}_i$ is the predicted value and $y_i$ is the original value for each instance $i$.
Focal¶
$$\textrm{FL} = -\alpha_{t}(1 - p_{t})^{\gamma}\log{p_{t}}$$$\gamma$ controls the shape of the curve. The higher the value of $\gamma$, the lower the loss for well-classified examples, so we could turn the attention of the model more towards ‘hard-to-classify examples. FL gives high weights to the rare class and small weights to the dominating or common class. These weights are referred to as $\alpha$.
Code¶
tf.keras.losses.BinaryCrossentropytfa.losses.SigmoidFocalCrossEntropy-->
Build Model¶
from keras_cv_attention_models import efficientnet
def build_model(model_name=CFG.model_name,
loss_name=CFG.loss,
dim=CFG.img_size,
compile_model=True,
include_top=False):
# Define backbone
base = getattr(efficientnet, model_name)(input_shape=(*dim,3),
pretrained='imagenet',
num_classes=0) # get base model (efficientnet), use imgnet weights
inp = base.inputs
x = base.output
x = tf.keras.layers.GlobalAveragePooling2D()(x) # use GAP to get pooling result form conv outputs
# Define 'necks' for each head
x_bowel = tf.keras.layers.Dense(32, activation='silu')(x)
x_extra = tf.keras.layers.Dense(32, activation='silu')(x)
x_liver = tf.keras.layers.Dense(32, activation='silu')(x)
x_kidney = tf.keras.layers.Dense(32, activation='silu')(x)
x_spleen = tf.keras.layers.Dense(32, activation='silu')(x)
# Define heads
out_bowel = tf.keras.layers.Dense(1, name='bowel', activation='sigmoid')(x_bowel) # use sigmoid to convert predictions to [0-1]
out_extra = tf.keras.layers.Dense(1, name='extra', activation='sigmoid')(x_extra) # use sigmoid to convert predictions to [0-1]
out_liver = tf.keras.layers.Dense(3, name='liver', activation='softmax')(x_liver) # use softmax for the liver head
out_kidney = tf.keras.layers.Dense(3, name='kidney', activation='softmax')(x_kidney) # use softmax for the kidney head
out_spleen = tf.keras.layers.Dense(3, name='spleen', activation='softmax')(x_spleen) # use softmax for the spleen head
# Combine outputs
out = [out_bowel, out_extra, out_liver, out_kidney, out_spleen]
# Create model
model = tf.keras.Model(inputs=inp, outputs=out)
if compile_model:
# optimizer
opt = tf.keras.optimizers.Adam(learning_rate=0.0001)
# loss
loss = {
'bowel':tf.keras.losses.BinaryCrossentropy(label_smoothing=0.05),
'extra':tf.keras.losses.BinaryCrossentropy(label_smoothing=0.05),
'liver':tf.keras.losses.CategoricalCrossentropy(label_smoothing=0.05),
'kidney':tf.keras.losses.CategoricalCrossentropy(label_smoothing=0.05),
'spleen':tf.keras.losses.CategoricalCrossentropy(label_smoothing=0.05),
}
# metric
metrics = {
'bowel':['accuracy'],
'extra':['accuracy'],
'liver':['accuracy'],
'kidney':['accuracy'],
'spleen':['accuracy'],
}
# compile
model.compile(optimizer=opt,
loss=loss,
metrics=metrics)
return model
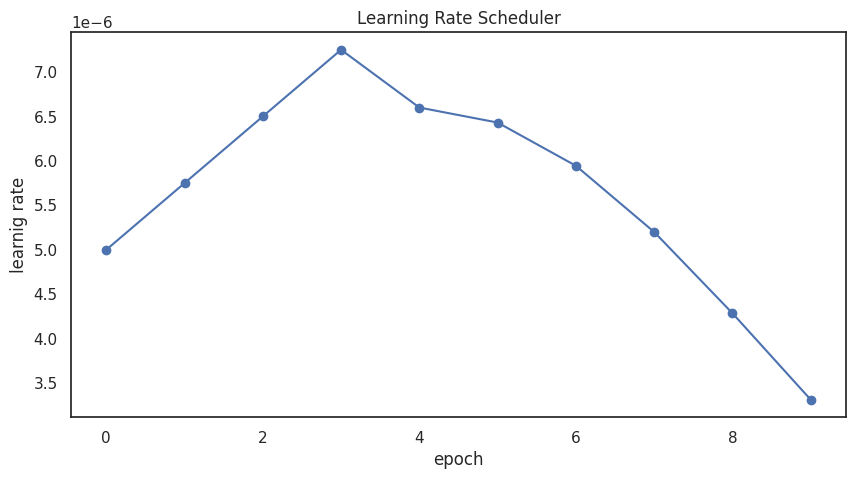
Learning-Rate Scheduler¶
def get_lr_callback(batch_size=8, plot=False):
lr_start = 0.000005
lr_max = 0.00000050 * REPLICAS * batch_size
lr_min = 0.000001
lr_ramp_ep = 4
lr_sus_ep = 0
lr_decay = 0.8
def lrfn(epoch):
if epoch < lr_ramp_ep:
lr = (lr_max - lr_start) / lr_ramp_ep * epoch + lr_start
elif epoch < lr_ramp_ep + lr_sus_ep:
lr = lr_max
elif CFG.scheduler=='exp':
lr = (lr_max - lr_min) * lr_decay**(epoch - lr_ramp_ep - lr_sus_ep) + lr_min
elif CFG.scheduler=='cosine':
decay_total_epochs = CFG.epochs - lr_ramp_ep - lr_sus_ep + 3
decay_epoch_index = epoch - lr_ramp_ep - lr_sus_ep
phase = math.pi * decay_epoch_index / decay_total_epochs
cosine_decay = 0.4 * (1 + math.cos(phase))
lr = (lr_max - lr_min) * cosine_decay + lr_min
return lr
if plot:
plt.figure(figsize=(10,5))
plt.plot(np.arange(CFG.epochs), [lrfn(epoch) for epoch in np.arange(CFG.epochs)], marker='o')
plt.xlabel('epoch'); plt.ylabel('learnig rate')
plt.title('Learning Rate Scheduler')
plt.show()
lr_callback = tf.keras.callbacks.LearningRateScheduler(lrfn, verbose=False)
return lr_callback
_=get_lr_callback(CFG.batch_size, plot=True )
import cv2
import pydicom
IMG_DIR = '/kaggle/working/'
def standardize_pixel_array(dcm: pydicom.dataset.FileDataset) -> np.ndarray:
# Correct DICOM pixel_array if PixelRepresentation == 1.
pixel_array = dcm.pixel_array
if dcm.PixelRepresentation == 1:
bit_shift = dcm.BitsAllocated - dcm.BitsStored
dtype = pixel_array.dtype
new_array = (pixel_array << bit_shift).astype(dtype) >> bit_shift
pixel_array = pydicom.pixel_data_handlers.util.apply_modality_lut(new_array, dcm)
return pixel_array
def read_xray(path, fix_monochrome = True):
dicom = pydicom.dcmread(path)
data = standardize_pixel_array(dicom)
data = data - np.min(data)
data = data / (np.max(data) + 1e-5)
if fix_monochrome and dicom.PhotometricInterpretation == "MONOCHROME1":
data = 1.0 - data
return data
def resize_and_save(file_path):
img = read_xray(file_path)
h, w = img.shape[:2] # orig hw
img = cv2.resize(img, (CFG.resize_dim, CFG.resize_dim), cv2.INTER_LINEAR)
img = (img * 255).astype(np.uint8)
sub_path = file_path.split("/",4)[-1].split('.dcm')[0] + '.png'
infos = sub_path.split('/')
sub_path = file_path.split("/",4)[-1].split('.dcm')[0] + '.png'
infos = sub_path.split('/')
pid = infos[-3]
sid = infos[-2]
iid = infos[-1]; iid = iid.replace('.png','')
new_path = os.path.join(IMG_DIR, sub_path)
os.makedirs(new_path.rsplit('/',1)[0], exist_ok=True)
cv2.imwrite(new_path, img)
return
Train Model¶
Cross-Validation: 5 fold
scores = []
for fold in np.arange(CFG.folds):
# Stop watch (start)
start_time = time.time()
# ignore not selected folds
if fold not in CFG.selected_folds:
continue
# train and valid dataframe
train_df = df[df["fold"]!=fold]
valid_df = df[df["fold"]==fold]
# get image_paths and labels
train_paths = train_df.image_path.values; train_labels = train_df[CFG.target_col].values.astype(np.float32)
valid_paths = valid_df.image_path.values; valid_labels = valid_df[CFG.target_col].values.astype(np.float32)
##test_paths = test_df.image_path.values
# shuffle train data
index = np.arange(len(train_df))
np.random.shuffle(index)
train_paths = train_paths[index]
train_labels = train_labels[index]
# min samples in debug mode
min_samples = CFG.batch_size*REPLICAS*2
# for debug model run on small portion
if CFG.debug:
train_paths = train_paths[:min_samples]; train_labels = train_labels[:min_samples]
valid_paths = valid_paths[:min_samples]; valid_labels = valid_labels[:min_samples]
# show message
print('#'*40); print('#### FOLD: ',fold)
print('#### IMAGE_SIZE: (%i, %i) | MODEL_NAME: %s | BATCH_SIZE: %i'%
(CFG.img_size[0],CFG.img_size[1],CFG.model_name,CFG.batch_size*REPLICAS))
# data stat
num_train = len(train_paths)
num_valid = len(valid_paths)
print('#### NUM_TRAIN: {:,} | NUM_VALID: {:,}'.format(num_train, num_valid))
# build model
K.clear_session()
with strategy.scope():
model = build_model(CFG.model_name, dim=CFG.img_size, compile_model=True)
# build dataset
cache = 1 if 'TPU' in CFG.device else 0
train_ds = build_dataset(train_paths, train_labels, cache=cache, batch_size=CFG.batch_size*REPLICAS,
repeat=True, shuffle=True, augment=CFG.augment)
val_ds = build_dataset(valid_paths, valid_labels, cache=cache, batch_size=CFG.batch_size*REPLICAS,
repeat=False, shuffle=False, augment=False)
print('#'*40)
# callbacks
callbacks = []
## save best model after each fold
sv = tf.keras.callbacks.ModelCheckpoint(
'fold-%i.h5'%fold, monitor='val_loss', verbose=CFG.verbose, save_best_only=True,
save_weights_only=False, mode='min', save_freq='epoch')
callbacks +=[sv]
## lr-scheduler
callbacks += [get_lr_callback(CFG.batch_size)]
# train
print('Training...')
history = model.fit(
train_ds,
epochs=CFG.epochs if not CFG.debug else 2,
callbacks = callbacks,
steps_per_epoch=len(train_paths)/CFG.batch_size//REPLICAS,
validation_data=val_ds,
verbose=CFG.verbose
)
# store best results
best_epoch = np.argmin(history.history['val_loss'])
best_loss = history.history['val_loss'][best_epoch]
best_acc_bowel = history.history['val_bowel_accuracy'][best_epoch]
best_acc_extra = history.history['val_extra_accuracy'][best_epoch]
best_acc_liver = history.history['val_liver_accuracy'][best_epoch]
best_acc_kidney = history.history['val_kidney_accuracy'][best_epoch]
best_acc_spleen = history.history['val_spleen_accuracy'][best_epoch]
# Find mean accuracy
best_acc = np.mean([best_acc_bowel, best_acc_extra,
best_acc_liver, best_acc_kidney, best_acc_spleen])
print(f'\n{"="*17} FOLD {fold} RESULTS {"="*17}')
print(f'>>>> BEST Loss : {best_loss:.3f}\n>>>> BEST Acc : {best_acc:.3f}\n>>>> BEST Epoch : {best_epoch}\n')
print('ORGAN Acc:')
print(f' >>>> {"Bowel".ljust(15)} : {best_acc_bowel:.3f}')
print(f' >>>> {"Extravasation".ljust(15)} : {best_acc_extra:.3f}')
print(f' >>>> {"Liver".ljust(15)} : {best_acc_liver:.3f}')
print(f' >>>> {"Kidney".ljust(15)} : {best_acc_kidney:.3f}')
print(f' >>>> {"Spleen".ljust(15)} : {best_acc_spleen:.3f}')
print(f'{"="*50}\n\n')
scores.append([best_loss, best_acc,
best_acc_bowel, best_acc_extra,
best_acc_liver, best_acc_kidney, best_acc_spleen])
#Stop watch (stop)
stop_time = time.time()
process_time = (stop_time - start_time) // 60
print(f"{process_time}[min]\n\n")
########################################
#### FOLD: 0
#### IMAGE_SIZE: (256, 256) | MODEL_NAME: EfficientNetV1B0 | BATCH_SIZE: 16
#### NUM_TRAIN: 9,817 | NUM_VALID: 2,212
Downloading data from https://github.com/leondgarse/keras_efficientnet_v2/releases/download/effnetv1_pretrained/efficientnetv1-b0-imagenet.h5
[Error] will not load weights, url not found or download failed: https://github.com/leondgarse/keras_efficientnet_v2/releases/download/effnetv1_pretrained/efficientnetv1-b0-imagenet.h5
########################################
Training...
================= FOLD 0 RESULTS =================
>>>> BEST Loss : 3.435
>>>> BEST Acc : 0.735
>>>> BEST Epoch : 8
ORGAN Acc:
>>>> Bowel : 0.622
>>>> Extravasation : 0.716
>>>> Liver : 0.881
>>>> Kidney : 0.694
>>>> Spleen : 0.763
==================================================
24.0[min]
########################################
#### FOLD: 1
#### IMAGE_SIZE: (256, 256) | MODEL_NAME: EfficientNetV1B0 | BATCH_SIZE: 16
#### NUM_TRAIN: 8,975 | NUM_VALID: 3,054
Downloading data from https://github.com/leondgarse/keras_efficientnet_v2/releases/download/effnetv1_pretrained/efficientnetv1-b0-imagenet.h5
[Error] will not load weights, url not found or download failed: https://github.com/leondgarse/keras_efficientnet_v2/releases/download/effnetv1_pretrained/efficientnetv1-b0-imagenet.h5
########################################
Training...
================= FOLD 1 RESULTS =================
>>>> BEST Loss : 3.291
>>>> BEST Acc : 0.751
>>>> BEST Epoch : 2
ORGAN Acc:
>>>> Bowel : 0.547
>>>> Extravasation : 0.716
>>>> Liver : 0.843
>>>> Kidney : 0.916
>>>> Spleen : 0.734
==================================================
22.0[min]
########################################
#### FOLD: 2
#### IMAGE_SIZE: (256, 256) | MODEL_NAME: EfficientNetV1B0 | BATCH_SIZE: 16
#### NUM_TRAIN: 9,012 | NUM_VALID: 3,017
Downloading data from https://github.com/leondgarse/keras_efficientnet_v2/releases/download/effnetv1_pretrained/efficientnetv1-b0-imagenet.h5
[Error] will not load weights, url not found or download failed: https://github.com/leondgarse/keras_efficientnet_v2/releases/download/effnetv1_pretrained/efficientnetv1-b0-imagenet.h5
########################################
Training...
================= FOLD 2 RESULTS =================
>>>> BEST Loss : 3.759
>>>> BEST Acc : 0.674
>>>> BEST Epoch : 0
ORGAN Acc:
>>>> Bowel : 0.461
>>>> Extravasation : 0.705
>>>> Liver : 0.870
>>>> Kidney : 0.854
>>>> Spleen : 0.477
==================================================
22.0[min]
Calculate OOF Score¶
# overall oof pF1
oof_loss, oof_acc, oof_acc_bowel, oof_acc_extra, oof_acc_liver, oof_acc_kidney, oof_acc_spleen = np.array(scores).mean(axis=0)
print(f'\n{"="*15} OVERALL OOF RESULTS {"="*15}')
print(f'>>>> OOF BEST Loss : {oof_loss:.3f}\n>>>> OOF BEST Acc : {oof_acc:.3f}\n')
print('ORGAN OOF Acc:')
print(f' >>>> {"Bowel".ljust(15)} : {oof_acc_bowel:.3f}')
print(f' >>>> {"Extravasation".ljust(15)} : {oof_acc_extra:.3f}')
print(f' >>>> {"Liver".ljust(15)} : {oof_acc_liver:.3f}')
print(f' >>>> {"Kidney".ljust(15)} : {oof_acc_kidney:.3f}')
print(f' >>>> {"Spleen".ljust(15)} : {oof_acc_spleen:.3f}')
print(f'{"="*50}\n')
=============== OVERALL OOF RESULTS ===============
>>>> OOF BEST Loss : 3.495
>>>> OOF BEST Acc : 0.720
ORGAN OOF Acc:
>>>> Bowel : 0.543
>>>> Extravasation : 0.712
>>>> Liver : 0.865
>>>> Kidney : 0.821
>>>> Spleen : 0.658
==================================================
Prepare Test dataset
#test_folder metadata !!!!NOTE THIS IS FOR TEST DATA !!!!
import itertools
#get patients
#patients = [patient for patient in os.listdir("/kaggle/input/rsna-2023-abdominal-trauma-detection/test_images") if os.path.isdir("/kaggle/input/rsna-2023-abdominal-trauma-detection/test_images/"+patient)]
patients = [patient for patient in os.listdir("/kaggle/input/rsna-2023-abdominal-trauma-detection/test_images")]
print("test_patient_id:", patients)
#get series
series = [os.listdir("/kaggle/input/rsna-2023-abdominal-trauma-detection/test_images/" + patient)[0] for patient in patients]
#series = list(itertools.chain.from_iterable(series)) #detouch double brackets
print("test_series_id:", series)
#get img
imgs = [os.listdir(f"/kaggle/input/rsna-2023-abdominal-trauma-detection/test_images/{patients[i]}/{series[i]}")[0] for i in range(len(patients))]
#imgs = list(itertools.chain.from_iterable(imgs)) #detouch double brackets
print("test_img_id:", imgs)
#make path
test_path=[f"/kaggle/input/rsna-2023-abdominal-trauma-detection/test_images/{patients[i]}/{series[i]}/{imgs[i]}" for i in range(len(patients))]
test_df = pd.DataFrame({
"patient_id": patients,
"series_id": series,
"image": imgs,
"path": test_path
})
test_df
test_patient_id: ['63706', '50046', '48843']
test_series_id: ['39279', '24574', '62825']
test_img_id: ['30.dcm', '30.dcm', '30.dcm']
| patient_id | series_id | image | path | |
|---|---|---|---|---|
| 0 | 63706 | 39279 | 30.dcm | /kaggle/input/rsna-2023-abdominal-trauma-detec... |
| 1 | 50046 | 24574 | 30.dcm | /kaggle/input/rsna-2023-abdominal-trauma-detec... |
| 2 | 48843 | 62825 | 30.dcm | /kaggle/input/rsna-2023-abdominal-trauma-detec... |
import gc
#%%time
from joblib import Parallel, delayed
file_paths = test_df.path
_ = Parallel(n_jobs=2,backend='threading')(delayed(resize_and_save)(file_path)\
for file_path in tqdm(file_paths, leave=True, position=0))
del _; gc.collect()
path_arr_img = [f"/kaggle/working/test_images/{patients[i]}/{series[i]}/{imgs[i]}".replace(".dcm", ".png") for i in range(len(patients))]
arr_df = pd.DataFrame(path_arr_img, columns=["arr_img_path"])
test_df = pd.concat([test_df, arr_df], axis=1)
test_df
100%|██████████| 3/3 [00:00<00:00, 1131.46it/s]
| patient_id | series_id | image | path | arr_img_path | |
|---|---|---|---|---|---|
| 0 | 63706 | 39279 | 30.dcm | /kaggle/input/rsna-2023-abdominal-trauma-detec... | /kaggle/working/test_images/63706/39279/30.png |
| 1 | 50046 | 24574 | 30.dcm | /kaggle/input/rsna-2023-abdominal-trauma-detec... | /kaggle/working/test_images/50046/24574/30.png |
| 2 | 48843 | 62825 | 30.dcm | /kaggle/input/rsna-2023-abdominal-trauma-detec... | /kaggle/working/test_images/48843/62825/30.png |
test_ds = build_dataset(test_df.arr_img_path,augment=False, repeat=False)
pred_results = model.predict(test_ds)
pred_results
1/1 [==============================] - 2s 2s/step
[array([[0.61479986],
[0.5274283 ],
[0.5548655 ]], dtype=float32),
array([[0.66972625],
[0.64082223],
[0.5338411 ]], dtype=float32),
array([[0.8420402 , 0.07981548, 0.07814431],
[0.83090025, 0.04297536, 0.12612435],
[0.810051 , 0.14346236, 0.04648664]], dtype=float32),
array([[0.8450608 , 0.10550442, 0.04943471],
[0.6726465 , 0.26745588, 0.05989753],
[0.7807357 , 0.19236377, 0.02690054]], dtype=float32),
array([[0.461854 , 0.19578502, 0.34236103],
[0.6382075 , 0.07609551, 0.28569704],
[0.7840603 , 0.04230442, 0.17363524]], dtype=float32)]
#patient_id
patient_df = test_df.patient_id
#binary_opposition
bowel_opposition_array = []
extravasation_opposition_array = []
for i in range(len(patients)):
bowel_opposition_array.append(1 - pred_results[0][i])
extravasation_opposition_array.append(1 - pred_results[1][i])
bowel_opposition_df = pd.DataFrame(bowel_opposition_array)
extravasation_opposition_df = pd.DataFrame(extravasation_opposition_array)
#binary
bowel_df = pd.DataFrame(pred_results[0])
extravasation_df = pd.DataFrame(pred_results[1])
#multi_value
kidney_df = pd.DataFrame(pred_results[2])
liver_df = pd.DataFrame(pred_results[3])
spleen_df = pd.DataFrame(pred_results[4])
columns = ["patient_id", "bowel_healthy", "bowel_injury", "extravasation_healthy", "extravasation_injury", "kidney_healthy", "kidney_low", "kidney_high", "liver_healthy", "liver_low", "liver_high", "spleen_healthy", "spleen_low", "spleen_high"]
pred_df = pd.concat([patient_df, bowel_df, bowel_opposition_df, extravasation_df, extravasation_opposition_df, kidney_df, liver_df, spleen_df],axis=1)
pred_df.columns = columns
pred_df.to_csv("/kaggle/working/submission.csv",index=False)




コメント・お問合せ
以下のツイートの『返信』にてお願いいたします